Trinucleotide repeat disorders (also known as trinucleotide repeat expansion disorders, triplet repeat expansion disorders or codon reiteration disorders) are a set of genetic disorders caused by trinucleotide repeats in certain genes exceeding the normal, stable, threshold, which differs per gene. The mutation is a subset of unstable microsatellite repeats that occur throughout all genomic sequences. If the repeat is present in a healthy gene, a dynamic mutation may increase the repeat count and result in a defective gene.
Trinucleotide repeat disorders are classified as a type of Non-Mendelian inheritance


 Subscribe in a reader
Subscribe in a reader
Since the early 90’s, a new class of molecular disease has been characterized based upon the presence of unstable and abnormal expansions of DNA-triplets (trinucleotides). The first triplet disease to be identified was fragile X syndrome that has since been mapped to the long arm of the X chromosome. At this point, there are from 230 to 4000 CGG repeats in the gene that causes fragile X syndrome in these patients, as compared with 60 to 230 repeats in carriers and 5 to 54 repeats in normal persons. The chromosomal instability resulting from this trinucleotide expansion presents clinically as mental retardation, distinctive facial features, and macroorchidism in males. The second, related DNA-triplet repeat disease, fragile X-E syndrome, was also identified on the X chromosome, but was found to be the result of an expanded GCC repeat. Identifying trinucleotide repeats as the basis of disease has brought clarity to our understanding of a complex set of inherited neurologic diseases.
As more repeat expansion diseases have been discovered, several categories have been established to group them based upon similar characteristics. Category 1 includes Huntington’s disease (HD) and the spinocerebellar ataxias that are caused by a CAG repeat expansion in a protein-coding portion of specific genes. Category 2 expansions tend to be more phenotypically diverse with heterogeneous expansions that are generally small in magnitude, but also found in the exons of genes. Category 3 includes fragile X syndrome, myotonic dystrophy, two of the spinocerebellar ataxias, juvenile myoclonic epilepsy, and Friedreich’s ataxia. These diseases are characterized by typically much larger repeat expansions than the first two groups, and the repeats are located outside of the protein-coding regions of the genes.
Currently, ten neurologic disorders are known to be caused by an increased number of CAG repeats that encode an expanded series of glutamine residues in otherwise unrelated proteins. During protein synthesis, the expanded CAG repeats are translated into a series of uninterrupted glutamine residues forming what is known as a polyglutamine tract. These disorders are characterized by autosomal dominant mode of inheritance (with the exception of spino-bulbar muscular atrophy which shows X-linked inheritance), midlife onset, a progressive course, and a correlation of the number of CAG repeats with the severity of disease and the age at onset. Family studies have also suggested that these diseases are associated with anticipation, the tendency for progressively earlier or more severe expression of the disease in successive generations. Although the causative genes are widely expressed in all of the known polyglutamine diseases, each disease displays an extremely selective pattern of neurodegeneration.
Symptoms
A common symptom of Polyq diseases is characterized by a progressive degeneration of nerve cells usually affecting people later in life. Although these diseases share the same repeated codon (CAG) and some symptoms, the repeats for the different polyglutamine diseases occur on different chromosomes.
Trinucleotide repeat disorders generally show genetic anticipation, where their severity increases with each successive generation that inherits them.
Trinucleotide repeat disorders are the result of extensive duplication of a single codon. In fact, the cause is trinucleotide expansion up to a repeat number above a certain threshold level.
Why three nucleotides?
An interesting question is why three nucleotides are expanded, rather than two or four or some other number. Dinucleotide repeats are a common feature of the genome in general, as are larger repeats (e.g. VNTRs - Variable Number Tandem Repeats). One possibility is that repeats that are not a multiple of three would not be viable. Trinucleotide repeat expansions tend to be near coding regions of the genome, and therefore repeats that are not multiples of three could cause frameshift mutations that would be deadly
The non-Polyq diseases do not share any specific symptoms and are unlike the Polyq diseases.
Trinucleotide repeat expansion
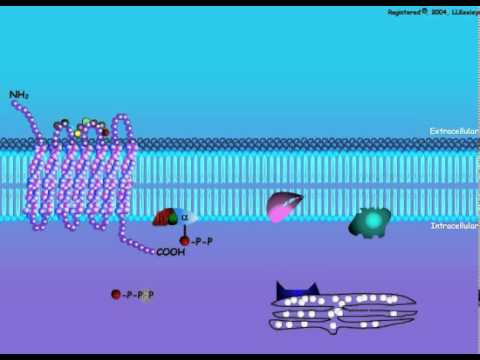
Trinucleotide repeat expansion, also known as triplet repeat expansion, is the DNA mutation responsible for causing any type of disorder categorized as a trinucleotide repeat disorder. Robert I. Richards and Grant R. Sutherland called these phenomena, in the framework of dynamical genetics, dynamic mutations.
Triplet expansion is caused by slippage during DNA replication. Due to the repetitive nature of the DNA sequence in these regions, 'loop out' structures may form during DNA replication while maintaining complementary base paring between the parent strand and daughter strand being synthesized. If the loop out structure is formed from sequence on the daughter strand this will result in an increase in the number of repeats. However if the loop out structure is formed on the parent strand a decrease in the number of repeats occurs. It appears that expansion of these repeats is more common than reduction. Generally the larger the expansion the more likely they are to cause disease or increase the severity of disease. This property results in the characteristic of anticipation seen in trinucleotide repeat disorders. Anticipation describes the tendency of age of onset to decrease and severity of symptoms to increase through successive generations of an affected family due to the expansion of these repeats.
In 2007, a team of scientists led by Ehud Shapiro at the Weizmann Institute of Science in Rehovot, Israel, proposed a new disease model to explain the progression of Huntington's Disease and similar trinucleotide repeat disorders. The team's computer simulations accurately predict age of onset and the way the disease will progress in an individual, based on the number of repeats of a genetic mutation.









.jpg)
.jpg)

.jpg)








