Jim Lockhart speaks with Anuradha Mittal of The Oakland Institute and Mark Des Marets of Northwest Resistance Against Genetic Engineering. Northwest Resistance Against Genetic Engineering www.nwrage.org From the webite SPEAKOUT! www.speakoutnow.org : Anuradha Mittal, a native of India, is an internationally renowned expert on trade, development, human rights, democracy, and agriculture issues. She is the founder and director of a policy think tank, The Oakland Institute, that works to ensure public participation and democratic debate on the most crucial economic and social policy issues that affect peoples' lives through nonpartisan research, analysis, and advocacy. Building on the identification of human rights as the foundation of global democracy, the Oakland Institute uses the rights-based approach to reframe the public debate and build an agenda for common action.
Prostate Removal Surgery
The word "laparoscopy" means to look inside the abdomen with a special camera or scope. Surgery performed with the aid of these cameras is known as "keyhole," "porthole," or "minimally invasive" surgery.
Traditional surgery requires a long incision (cut) down the center of the abdomen and a lengthy recovery period. Laparoscopic surgery eliminates the need for this large incision. As a result, you may experience less pain and scarring after surgery, more rapid recovery, and less risk of infection.
For laparoscopic prostate surgery, this technique requires five small (5-10 millimeters) incisions (or "portholes") -- one just below the belly button and two each on both sides of the lower abdomen. Carbon dioxide is passed into the abdominal cavity through a small tube placed into the incision below the belly button. This gas lifts the abdominal wall to give the surgeon a better view of the abdominal cavity once the laparoscope is in place. The surgeon is then guided by the laparoscope, which transmits a picture of the prostate onto a video monitor.
Laparoscopy is a relatively new technique for prostate removal, but it looks promising. Men who undergo this technique have less blood loss, less need for pain medicine, shorter hospital stays, quicker return to regular diet and activities, early removal of urethral catheters (tubes inserted through the penis to drain urine from the bladder), and a quicker recovery. Laparoscopy also appears to treat the cancer as well as conventional "open" procedures that are performed with a large incision.
What are the advantages of laparoscopy?
As is the case with other minimally invasive procedures, laparoscopic prostate removal has significant advantages over traditional "open" surgery:
Laparoscopy can shorten your hospital stay to one or two days. About 50 percent of men are discharged one day after surgery. (The length of stay depends on how quickly you recover and the extent of the surgery.)
There is significantly less bleeding during the operation.
You are less likely to need prescription painkillers after you leave the hospital. Patients generally need nothing more than Tylenol.At your follow-up appointment one week after surgery, the tube — or catheter — draining your bladder will be removed if there are no signs of other problems. Occasionally, the catheter must remain in place for another week, as is routinely the case following conventional "open" surgery.
About 90 percent of patients can return to work or resume full activity in only two to three weeks.
Am I eligible for this surgery?You are eligible if you have cancer that has not spread outside the prostate and is not very aggressive, as well as a PSA blood test less than 10. You are not eligible if you have had previous open or laparoscopic pelvic surgery, even for another reason, or a history of hormone treatment called LH-RH agonist (luteinizing hormone-releasing hormone), which reduces the size of the prostate tumor.
What are the side effects?
Medical research so far has shown the frequency of incontinence and impotence to be similar between minimally invasive surgery and open surgery -- with men usually returning to normal urinary function within three months for both types of surgeries. Both types of surgeries also have similar rates of incontinence.
Because this technique is nerve-sparing, postoperative sexual potency rate should be comparable to that of conventional open surgery. However, it is important to note that minimally invasive prostate surgery has not been in use long enough to truly assess whether or not it leads to higher rates of potency. But early results are promising.
How do I prepare for surgery?
Your surgeon will meet with you to answer any questions you might have. You will be asked questions about your health history, and a general physical examination will be performed. If your intestine requires cleaning, you will be given a prescription for a laxative medicine to take the evening before the surgery.
All patients are generally asked to provide a blood sample. Depending on your age and general health, you might also have an EKG (electrocardiogram), a chest X-ray, lung function tests, or other tests.
Finally, you will meet with an anesthesiologist who will discuss the type of anesthesia you will be given for surgery. You will also learn about pain control after the operation, which might include a PCA (patient controlled analgesia) pump.
What happens during surgery?
Your surgeon will place a small needle just below your belly button and insert the needle into your abdominal cavity. The needle is connected to a small tube, and carbon dioxide is passed into the abdominal cavity. This gas lifts the abdominal wall to give the surgeon a better view of the abdominal cavity once the laparoscope is in place. The surgeon is then guided by the laparoscope, which transmits a picture of the prostate onto a video monitor.
Next, a small incision will be made near your belly button. The laparoscope is placed through this incision and is connected to a video camera. The image your surgeon sees in the laparoscope is projected onto video monitors placed near the operating table.
Before starting the surgery, the surgeon will take a thorough look at your abdominal cavity to make sure the laparoscopy procedure will be safe for you. If the surgeon determines that the procedure will not be safe for you because of the presence of scar tissue, infection ,or abdominal disease, the procedure will not be continued.
If the surgeon decides the surgery can be safely performed, additional small incisions will be made, giving your surgeon access to the abdominal cavity. If necessary, one of these small incisions might be enlarged to remove the pelvic lymph nodes.
What happens after surgery?
You can expect to follow a liquid diet and you gradually will be able to eat solid foods. When you go home, you will follow a soft diet, which generally means no raw fruits or vegetables. A dietitian can provide more specific dietary guidelines.
Nausea and vomiting are common and occur because the intestines are temporarily disabled during anesthesia and surgery. Your doctor can prescribe medicines to relieve these symptoms, which will improve a few days after surgery.
You will be encouraged to get out of bed and walk as much as possible, starting the first day after surgery. You should steadily increase your activity after you go home. For six weeks after surgery, you should not lift or push anything over 30 pounds, and do not do abdominal exercises such as sit-ups.
Roger Kornberg lecture
Roger David Kornberg (born April 24, 1947) is an American biochemist and professor of structural biology at Stanford University School of Medicine.
Kornberg was awarded the Nobel Prize in Chemistry in 2006 for his studies of the process by which genetic information from DNA is copied to RNA, "the molecular basis of eukaryotic transcription." His father, Arthur Kornberg, who was also a professor at Stanford University, was awarded the Nobel Prize in Physiology or Medicine in 1959.
Part 1
part 2
part 3
part4
Part5
Part 6
Part 7
Scientific Discoveries and Nobel Prize
All organisms are controlled by their genes, which are coded by DNA, which is copied to RNA, which creates proteins, which are sequences of amino acids. DNA resides in the nucleus. When a cell expresses a gene, it copies (transcribes) that gene's DNA sequence onto a messenger RNA (mRNA) sequence. mRNA is transported out of the nucleus to ribosomes. The ribosomes read the mRNA and translate the code into the right amino acid sequence to make that gene's protein.
The DNA is transcribed to mRNA by an enzyme, RNA polymerase II, with the help of many other proteins. Using yeast, Kornberg identified the role of RNA polymerase II and other proteins in transcribing DNA, and he created three-dimensional images of the protein cluster using X-ray crystallography. Polymerase II is used by all organisms with nuclei, including humans, to transcribe DNA.
Kornberg and his research group have made several fundamental discoveries concerning the mechanisms and regulation of eukaryotic transcription. While a postdoctoral fellow working with Aaron Klug and Francis Crick at the MRC in the 1970s, Kornberg discovered the nucleosome as the basic protein complex packaging chromosomal DNA in the nucleus of eukaryotic cells (chromosomal DNA is often termed "Chromatin" when it is bound to proteins in this manner, reflecting Walther Flemming's discovery that certain structures within the cell nucleus would absorb dyes and become visible under a microscope).Within the nucleosome, Kornberg found that roughly 200 bp of DNA are wrapped around an octamer of histone proteins.
Kornberg's research group at Stanford later succeeded in the development of a faithful transcription system from baker's yeast, a simple unicellular eukaryote, which they then used to isolate in a purified form all of the several dozen proteins required for the transcription process. Through the work of Kornberg and others, it has become clear that these protein components are remarkably conserved across the full spectrum of eukaryotes, from yeast to human cells.
Using this system, Kornberg made the major discovery that transmission of gene regulatory signals to the RNA polymerase machinery is accomplished by an additional protein complex that they dubbed Mediator. As noted by the Nobel Prize committee, "the great complexity of eukaryotic organisms is actually enabled by the fine interplay between tissue-specific substances, enhancers in the DNA and Mediator. The discovery of Mediator is therefore a true milestone in the understanding of the transcription process."
At the same as Kornberg was pursuing these biochemical studies of the transcription process, he devoted two decades to the development of methods to visualize the atomic structure of RNA polymerase and its associated protein components. Initially, Kornberg took advantage of expertise with lipid membranes gained from his graduate studies to devise a technique for the formation of two-dimensional protein crystals on lipid bilayers. These 2D crystals could then be analyzed using electron microscopy to derive low-resolution images of the protein's structure. Eventually, Kornberg was able to use X-ray crystallography to solve the 3-dimensional structure of RNA polymerase at atomic resolution. The structure of RNA polymerase obtained by Kornberg is the most complex protein structure solved to date. He has recently extended these studies to obtain structural images of RNA polymerase associated with accessory proteins.[8]Through these studies, Kornberg has created an actual picture of how transcription works at a molecular level. According to the Nobel Prize committee, "the truly revolutionary aspect of the picture Kornberg has created is that it captures the process of transcription in full flow. What we see is an RNA-strand being constructed, and hence the exact positions of the DNA, polymerase and RNA during this process."
In 1959, Roger Kornberg's father, Arthur Kornberg, received the Nobel Prize in Physiology or Medicine for studies of how genetic information is transferred from one DNA molecule to another in a process called DNA replication. Specifically, Arthur Kornberg isolated the first enzyme capable of synthesizing DNA, bacterial DNA polymerase I, which was then the first known enzyme to take its instructions from a template, thus ensuring the conservation of genetic information during cellular growth and division. Roger Kornberg's younger brother, Thomas Bill Kornberg, discovered DNA polymerases II and III in 1970 and is now a geneticist at the University of California, San Francisco. All three Kornbergs have thus worked to understand how genetic information is put to use in cells. Roger and Arthur Kornberg are the sixth father-son pair to win Nobel Prizes.
Roger D. Kornberg. (2009, January 21). In Wikipedia, The Free Encyclopedia. Retrieved 05:03, February 24, 2009, from http://en.wikipedia.org/w/index.php?title=Roger_D._Kornberg&oldid=265517564
Roger D. Kornberg. (2009, January 21). In Wikipedia, The Free Encyclopedia. Retrieved 05:03, February 24, 2009, from http://en.wikipedia.org/w/index.php?title=Roger_D._Kornberg&oldid=265517564
Return to the RNAi World: Rethinking Gene Expression, Evolution and Medicine
Dr.Cameron Mello was awarded the 2006 Nobel prize for Medicine , along with colleague Dr. Andrew Z. Fire, for the discovery of RNA interference. In this video, he discusses his research and his experiences as a scientist and recent Nobel Prize
Craig Cameron Mello (born October 18, 1960) is an American biologist and Professor of Molecular Medicine at the University of Massachusetts Medical School in Worcester, Massachusetts. He was awarded the 2006 Nobel Prize for Physiology or Medicine, along with Andrew Z. Fire, for the discovery of RNA interference. This research was conducted at the University of Massachusetts Medical School and published in 1998. Mello has been a Howard Hughes Medical Institute investigator since 2000.
About Speaker
Mello and Fire received the Nobel Prize for work that began in 1998, when Mello and Fire along with their colleagues (SiQun Xu, Mary Montgomery, Stephen Kostas, and Sam Driver) published a paper [3] in the journal Nature detailing how tiny snippets of RNA fool the cell into destroying the gene's messenger RNA (mRNA) before it can produce a protein - effectively shutting specific genes down.
Nobel Prize for Medecine 2008
Nobel prize for medicine 2008 was Shared between Harald zur Hausen "for his discovery of human papilloma viruses causing cervical cancer" and to Françoise Barré-Sinoussi and Luc Montagnier for their discovery of human immunodeficiency virus.
Harald zur Hausen (born March 11, 1936) is a German virologist and professor emeritus. He has done research on cancer of the cervix, where he discovered the role of papilloma viruses,
Discovery of human papilloma virus causing cervical cancer
Against the prevailing view during the 1970s, Harald zur Hausen postulated a role for human papilloma virus (HPV) in cervical cancer. He assumed that the tumour cells, if they contained an oncogenic virus, should harbour viral DNA integrated into their genomes. The HPV genes promoting cell proliferation should therefore be detectable by specifically searching tumour cells for such viral DNA. Harald zur Hausen pursued this idea for over 10 years by searching for different HPV types, a search made difficult by the fact that only parts of the viral DNA were integrated into the host genome. He found novel HPV-DNA in cervix cancer biopsies, and thus discovered the new, tumourigenic HPV16 type in 1983. In 1984, he cloned HPV16 and 18 from patients with cervical cancer. The HPV types 16 and 18 were consistently found in about 70% of cervical cancer biopsies throughout the world.
Françoise Barré-Sinoussi
Françoise Barré-Sinoussi (born 30 July 1947) is a French virologist and director of the Unité de Régulation des Infections Rétrovirales at the Institut Pasteur in Paris, France. Born in Paris, France, Barré-Sinoussi performed some of the fundamental work in the identification of the human immunodeficiency virus (HIV) as the cause of AIDS
Luc Montagnier (born August 18, 1932) is a French virologist and joint recipient with Françoise Barré-Sinoussi and Harald zur Hausen of the 2008 Nobel Prize in Physiology or Medicine.
Discovery of HIV
Following medical reports of a novel immunodeficiency syndrome in 1981, the search for a causative agent was on. Françoise Barré-Sinoussi and Luc Montagnier isolated and cultured lymph node cells from patients that had swollen lymph nodes characteristic of the early stage of acquired immune deficiency. They detected activity of the retroviral enzyme reverse transcriptase, a direct sign of retrovirus replication. They also found retroviral particles budding from the infected cells. Isolated virus infected and killed lymphocytes from both diseased and healthy donors, and reacted with antibodies from infected patients. In contrast to previously characterized human oncogenic retroviruses, the novel retrovirus they had discovered, now known as human immunodeficiency virus (HIV), did not induce uncontrolled cell growth. Instead, the virus required cell activation for replication and mediated cell fusion of T lymphocytes. This partly explained how HIV impairs the immune system since the T cells are essential for immune defence. By 1984, Barré-Sinoussi and Montagnier had obtained several isolates of the novel human retrovirus, which they identified as a lentivirus, from sexually infected individuals, haemophiliacs, mother to infant transmissions and transfused patients. The significance of their achievements should be viewed in the context of a global ubiquitous epidemic affecting close to 1% of the population.
Importance of the HIV discoverySoon after the discovery of the virus, several groups contributed to the definitive demonstration of HIV as the cause of acquired human immunodeficiency syndrome (AIDS). Barré-Sinoussi and Montagnier's discovery made rapid cloning of the HIV-1 genome possible. This has allowed identification of important details in its replication cycle and how the virus interacts with its host. Furthermore, it led to development of methods to diagnose infected patients and to screen blood products, which has limited the spread of the pandemic. The unprecedented development of several classes of new antiviral drugs is also a result of knowledge of the details of the viral replication cycle. The combination of prevention and treatment has substantially decreased spread of the disease and dramatically increased life expectancy among treated patients. The cloning of HIV enabled studies of its origin and evolution. The virus was probably passed to humans from chimpanzees in West Africa early in the 20th century, but it is still unclear why the epidemic spread so dramatically from 1970 and onwards.
Identification of virus−host interactions has provided information on how HIV evades the host’s immune system by impairing lymphocyte function, by constantly changing and by hiding its genome in the host lymphocyte DNA, making its eradication in the infected host difficult even after long-term antiviral treatment. Extensive knowledge about these unique viral host interactions has, however, generated results that can provide ideas for future vaccine development as well as for therapeutic approaches targeting viral latency.
HIV has generated a novel pandemic. Never before has science and medicine been so quick to discover, identify the origin and provide treatment for a new disease entity. Successful anti-retroviral therapy results in life expectancies for persons with HIV infection now reaching levels similar to those of uninfected people
What is Ovulation

Ovulation is the process in the menstrual cycle by which a mature ovarian follicle ruptures and discharges an ovum (also known as an oocyte, female gamete, or casually, an egg) that participates in reproduction. Ovulation also occurs in the estrous cycle of other female mammals, which differs in many fundamental ways from the menstrual cycle.
The process of ovulation is controlled by the hypothalamus of the brain and through the release of hormones secreted in the anterior lobe of the pituitary gland, (Luteinizing hormone (LH) and Follicle-stimulating hormone (FSH)). In the follicular (pre-ovulatory) phase of the menstrual cycle, the ovarian follicle will undergo a series of transformations called cumulus expansion, this is stimulated by the secretion of FSH. After this is done, a hole called the stigma will form in the follicle, and the ovum will leave the follicle through this hole. Ovulation is triggered by a spike in the amount of FSH and LH released from the pituitary gland. During the luteal (post-ovulatory) phase, the ovum will travel through the fallopian tubes toward the uterus. If fertilized by a sperm, it may perform implantation there 6-12 days later.
Overview
In humans, the few days near ovulation constitute the fertile phase. The average time of ovulation is the fourteenth day of an average length (twenty-eight day) menstrual cycle. It is normal for the day of ovulation to vary from the average, with ovulation anywhere between the tenth and nineteenth day being common.
Cycle length alone is not a reliable indicator of the day of ovulation. While in general an earlier ovulation will result in a shorter menstrual cycle, and vice versa, the luteal (post-ovulatory) phase of the menstrual cycle may vary by up to a week between women.
OSTEO ARTHIRITIS

Osteoarthritis , is a group of diseases and mechanical abnormalities entailing painful degradation of joints, including cartilage and the subchondral bone next to it. Clinical symptoms of OA may include joint pain, tenderness, stiffness, inflammation, and creaking of joints. In OA, a variety of potential forces -- hereditary, developmental, metabolic, and mechanical -- may initiate processes leading to loss of cartilage -- a strong protein matrix that lubricates and cushions the joints. As the body struggles to contain ongoing damage, immune and regrowth process can accelerate damage. When bone surfaces become less well protected by cartilage, subchondral bone may be exposed and damaged, with regrowth leading to a proliferation of ivory-like, dense, reactive bone in central areas of cartilage loss, a process called eburnation. The patient increasingly experiences pain upon weight bearing, including walking and standing. Due to decreased movement because of the pain, regional muscles may atrophy, and ligaments may become more lax. OA is the most common form of arthritis
Implications of Synthetic Biology

Synthetic biology is a new area of biological research that combines science and engineering in order to design and build ("synthesize") novel biological functions and systems
There’s no mistaking Drew Endy’s profession: “I like to make things -- that’s what I do.” From his engineer’s perspective, the slow and painful methods of bioengineering demand a solution. Endy hopes to refine the tools necessary to move the field forward. “We’re going from looking at the living world as only coming from nature, to a subset of the living world being produced by engineers who design and build hopefully useful living artifacts according to our specifications,” says Endy.
Thirty years ago, scientists figured out how to use enzymes to cut and paste genetic material, leading to recombinant DNA technology. But the techniques involved are painfully slow, requiring very specific physical materials and “know-how via the guild-like structure of biology.” Endy points to methods coming on line that will make it easier to design and build biological systems.
One is DNA synthesis, in which a machine fed information and sugars generates a physical piece of DNA. It reminds Endy of the “matter compilers” seen on Star Trek, where “food materializes from a cubby in the wall.” This technique will allow the economical production of long sequences of DNA. Another key ingredient in bioengineering will be the development of standards for making and measuring DNA, in the same way that machining hardware came to be governed by common standards in the 19th century. Endy also suggests that biotechnology will be increasingly informed by useful abstraction, so that scientists will manipulate raw materials less and refined and repackaged materials more, in order to make new things simply and more reliably. These advances will also enable bioengineers to “be experts in our own domains without having to be masters of everything.”
But as bioengineering becomes easier, and “people start to engineer biology,” we’ll need to worry about new issues, says Endy: Will people synthesize pathogens from scratch? Will groups pool knowledge legally? Will there be accreditation and oversight of those who create biological systems?
About the Speaker
Andrew Endy
Cabot Assistant Professor of Biological EngineeringDrew Endy earned degrees in civil, environmental, and biochemical engineering at Lehigh and Dartmouth. He studied genetics & microbiology as a postdoc at U.T. Austin and U.W. Madison. From 1998 through 2001 he helped to start the Molecular Sciences Institute, an independent not-for-profit biological research lab in Berkeley, CA.
In 2002, Endy started a group as a fellow in the Department of Biology and the Biological Engineering Division at MIT. He joined the MIT faculty in 2004. Endy co-founded the MIT Synthetic Biology working group and the Registry of Standard Biological Parts, and organized the First International Conference on Synthetic Biology. With colleagues he taught the 2003 and 2004 MIT Synthetic Biology labs that led to the organization of iGEM, the international Genetically Engineered Machine competiton.
In 2004 Endy co-founded Codon Devices, Inc., a venture-funded startup that is working to develop next-generation DNA synthesis technology. In 2005 Endy co-founded the BioBricks Foundation, a not-for-profit organization that is working to develop legal and economic strategies needed to support open biotechnology.
Metastasis Lecture
No diagnosis of cancer is welcome, but some scenarios are more dreaded than others. Richard Hynes discusses what happens “when cells in the primary tumor lose their sense of address and wander off to places they’re not supposed to go.” His talk lays out the process of invasion, by which the cancer spreads into tissues adjacent to the tumor, and that of metastasis, where the cancer disseminates to distant sites.
Hynes describes the transitions a cancer undergoes as it spreads. He explains how tissue in our bodies is made of sheets of epithelial cells that are carefully arranged on a “basement membrane” by a series of adhesion receptors. These receptors, if functioning properly, don’t usually allow the cells to go anywhere. When a cell becomes tumorigenic, it loses some adhesion, and then if it becomes more damaged “wanders off into the underlying tissue.” This is called invasion. Hynes and other researchers are looking at the molecules responsible for cells’ adhesive qualities, and at the mutations in genes that trigger a loss of adhesion. Some of these processes are part of normal development, but occasionally, a “switch gets thrown in cells that should have stayed epithelial” and they become migratory instead.
Once on the move, cancer cells “need plumbing to grow,” says Hynes. Tumors recruit blood vessels to feed them and remove waste, and they can also exploit the body’s white blood cells and platelets to promote their own growth. Hynes describes “cross talk between tumor cells and cells in bone,” where the “two cells get together in evil combination to damage the bone and enhance the growth of metastases.” Scientists have discovered “a lot of different mechanisms by which metastatic cells learn new tricks and suborn the mechanism of the host to get them where they’re going.” Hynes finds such insidious workings an “appealing thing, since these alterations offer opportunities for therapies.” Researchers can tinker with circuits between cells, restore growth suppression and interfere with blood vessel recruitment. It’s “a complex problem,” says Hynes, but there are “lots of ways to get at this.”
About the Speaker
Richard O. Hynes PhD
Daniel K. Ludwig Professor for Cancer Research, Department of Biology Investigator, Howard Hughes Medical InstituteRichard Hynes received his B.A. in biochemistry from the University of Cambridge, U.K., and his Ph.D. in biology from MIT. After postdoctoral work at the Imperial Cancer Research Fund in London, where he initiated his work on cell adhesion, he returned to MIT as a faculty member.
Hynes is a fellow of the Royal Society of London, the American Academy of Arts and Sciences, and the American Association for the Advancement of Science, and a member of the National Academy of Sciences and the Institute of Medicine. He has received the Gairdner Foundation International Award for achievement in medical science and recently served as president of the American Society for Cell Biology.
New Lessons in Cancer Research

Cancer is a conniving enemy. Try to kill it off through surgery or chemotherapy, and it finds a way to sneak back in. Jacqueline Lees tells an engaged Soap Box audience what insights and tools research now offers in the longstanding battle against this relentless disease.
Big gains have come from molecular study of tumors at different stages, Lees says. It often takes many years for a cancerous cell to develop into a dangerous tumor, one that can yield metastases. There might be six phases of development over 15 years in a cancer’s evolution, and scientists have formed a good understanding of what these different lesions look like in various cancers, and how they behave. Lees calls this process “actually a beautiful example of evolution,” since the cell that mutates and begins to divide uncontrollably evolves to become more successful relative to other cells in the tissue.
Other research focuses on the genetic basis of cancers. Two “flavors” of genes appear responsible for provoking cancerous changes in cells: oncogenes and tumor suppressor genes. It may be possible to intervene along the genetic pathways underlying cancer growth, says Lees. Her own work, involving mutant mice and zebrafish, hopes to identify the mechanisms involved in specific kinds of tumors, and to figure out ways of inhibiting cancer cell growth. Understanding the nature of specific cancers might help prevent treating people with chemical agents that don’t work for their kind of cancer, and that actually increase their tumor’s growth.
With the advent of fast and inexpensive genetic screens, it may soon be possible to determine whether each of us carries genes that predispose us toward certain kinds of cancers. But Lees questions the universal adoption of DNA testing, not just because of privacy concerns, but because there may very well be no known cure if a predisposition to disease is found. “If we sequenced every baby, and said you’re highly predisposed to a cancer, and there’s nothing we can do, would that be information people want to have?” Lees wonders. “If we could find a rapid way to sequence small subsets of genome, identify people with high risk and we could treat them if we knew they had those diseases, there’d be an argument for that, much as we do testing for diseases where we know can intervene if find children carrying them,” says Lees.
About the Speaker
Jacqueline Lees SM '86, PhD '90
Associate DirectorMIT Center for Cancer Research;
Professor of Biology
Jacqueline Lees' research is focused on identifying the proteins and pathways that play a key role in tumorigenicity and establishing the mechanism of their action in both normal and tumor cells. Her lab uses a combination of molecular and cellular analyses, mutant mouse models and genetic screens in zebrafish.
Lees received her Ph.D. in 1990 from the University of London.
Computational and Systems Approaches to Cancer

Early on in his lecture, Michael Yaffe serves up an amazing fact: If the distance between each DNA base pair were one foot apart, then each time a cell divided, it would have to copy 568 thousand miles of DNA. This, says Yaffe, is enough to go around the circumference of the earth more than 22 times. What’s more, the cell has to copy its DNA with no errors. “I don’t know (if) civil engineers ... could make 10 miles of road without making single error,” says Yaffe.
Yaffe’s in the business of exploring and mathematically mapping the elaborate signal pathways inside cells that sense broken DNA and coordinate damage response. While studying one such process, cell death in the colon, Yaffe found that the traditional biochemistry approach -- picking one molecule, one stimulus and one readout-- doesn’t work. “It’s like the blind man feeling the elephant’s tail, and saying it’s a long, thin animal.” Yaffe learned that one signal may activate a series of proteins, triggering an amplification loop. A slight change might yield a “whopping response.”
In the 12 hours a cell takes to copy its DNA to create two daughter cells, “it goes to great pains to make sure everything is done correctly. It initiates checkpoints, like border crossings.” Because everyday life exposes DNA to all kinds of damage, cells have evolved “an elaborate surveillance mechanism” to “blow the whistle, signal repair, and recruit repair machinery,” or if damage is too great, essentially commit suicide. If something goes wrong with this mechanism at crucial times during cell division, cancer frequently results.
Just as engineers test integrated circuits at a variety of points, Yaffe came up with a method of testing cell signaling with a variety of proteins. His team came up with 7,000 signaling measurements in 760 dimensions, and 1,400 signal responses. But this data-heavy model for predicting which molecules lead to cell death didn’t satisfy Yaffe. With additional mathematical sleight of hand, Yaffe’s group boiled down the cell signaling measurements to what Yaffe calls two “canonical super axes”: “a global measure of cell stress and death, and another of survival signaling.” He hopes to use this slimmed-down model to think about drugs targeting cancer and inflammation.
About the Speaker
Michael Yaffe
Howard S. and Linda B. Stern Associate Professor of Biology, Department of Biology and the Center for Cancer Research
Michael Yaffe received his Ph.D. in Biophysical Chemistry in 1987, and an M.D. in 1989, both from Case Western Reserve University. Before MIT, he served as a surgeon in teaching hospitals in Cleveland and the Boston area, including the Harvard Medical School.
He received multiple teaching awards from University Hospitals of Cleveland, and earned the 1998 Howard Hughes Physician Scientist Award, and the 1999 Burroughs Wellcome Career Development Award.
Cancer Research in the Genomic Era

Eric Lander likens the current age of biological discovery to the days of great ocean-going exploration. After the world was mapped, no one could imagine what it was like to live “before you knew what would happen if you sailed west.” Following the current revolution in biology, we “won’t be able to imagine what science was like...” This transformation, claims Lander, will be complete in the next decade or so. “MIT students in 2020 will look back with a mixture of amusement and horror at the late 20th century and say, ‘Imagine, people spent years looking for the gene for something.’”
Lander views biology as a vast library that will soon contain information not just about the DNA sequences of species, but ‘volumes’ on individuals, tissues, and cells. With great effort, researchers deciphered the secrets of chromosomes, the double helix, and more recently, the human genome and that of other species. But progress in such discoveries is now moving at a much faster clip due to high-speed computing and the Internet. MIT currently sequences ¼ million pieces of DNA per day, says Lander. He projects this pace will quicken by 20 fold in the next several years.
Fortified by this progress, Lander has compiled an ambitious ‘to-do list:’ identifying “everything that matters” in the human genome, from proteins to the things that control genes; knowing all human genetic variation in the population; knowing how to recognize when a cell “is thinking of one thing or another” based on how genes are turned on or off; knowing all the mechanisms that cause cancer and how to modulate all the genes.
Astonishingly, he says, “This is not the to-do list of the next century, but the next decade.” Lander is confident that researchers will in the not-distant future generate a catalog of the unique genetic signatures associated with “different flavors” of a type of cancer. Scientists will find patterns in diseases, genes and drug responses, and eventually assemble a list of all the genetic variants in the human genome that put individuals at risk for different diseases. These various gene databases will serve “as foundational information for biology for centuries to come,” concludes Lander.
About Speaker
Eric Lander was a world leader of the international Human Genome Project, the effort to map the blueprint for a human being. Today, Lander is using the knowledge of the human genome to tackle the fundamental issue of medicine: to find the causes of disease.
Lander received his Ph.D. in mathematics from Oxford in 1981, as a Rhodes Scholar. He joined Whitehead Institute in 1986 and founded the Whitehead Institute/MIT Center for Genome Research in 1990. Lander became the founding director of the newly created Broad Institute in 2003.
Lander is a member of the U.S. National Academy of Sciences, and U.S. Institute of Medicine. He was a MacArthur Fellow (1987-1992), and earned the Woodrow Wilson Prize from Princeton University(1998); the Baker Memorial Award for Undergraduate Teaching at MIT (1992); the City of Medicine Prize (2001); and the Gairdner International Prize (2002).
Engineering a New Attack on Disease

Out of a world population of 6 billion, 57 million people die each year. And while we have gained 20 years in life expectancy since World War 2, diseases like HIV have taken a toll on morbidity in many developing nations. But according to Rick Young, “the global disease burden is much larger than the number of deaths.” Countless millions suffer from cardiopulmonary diseases, cancer, and malaria, to name but a few, at a nearly incalculable cost to their families and society. Young’s mission is to attack the problem of global disease at the genetic level: he’s hunting for specific proteins that can turn the genetic machinery of diseases on, or off. These “gene regulators” can be knocked out of whack by a virus like HIV or by a mutation that results in a disease like mature onset diabetes. Young’s group has developed a DNA microarray technology that helps them link gene regulators to their corresponding genes. They’ve worked out the connections in yeast, and they’re targeting the human genome next. Young’s ultimate goal: “By continuing to focus on your 2000 gene regulators, we could eventually develop great insights into how organ systems work… (And) in all instances where disease is associated with misregulation, we could develop new strategies for drug development based on that.”
About the Speaker
Richard A. Young Member, Whitehead Institute Professor of Biology, MIT
Richard A. Young is a pioneer in gene transcription, the process by which cells read and interpret the genetic instructions embedded in DNA. His lab’s achievements include novel AIDS vaccine candidates and new approaches to drug-resistant tuberculosis.
Young received his Ph.D. in molecular biophysics and biochemistry from Yale University in 1979. He has been director of the National Cooperative Vaccine Development Group for AIDS at the Whitehead Institute and has served on several international committees for the World Health Organization. He received the Burroughs Wellcome Scholar Award in 1987 and the Chiron Corporation Biotechnology Research Award from the American Society for Microbiology in 1994. Young was elected a fellow of the American Academy of Microbiology in 1994, and a charter fellow of the Molecular Medicine Society in 1995.
Vitamin D: The New Old Natural Wonder Drug

Professor Hector F. DeLuca's laboratory has been devoted to the understanding of metabolism and mechanism of action of vitamins A and D. Sponsored by Wisconsin Academy of Sciences, Arts and Letters in partnership with the Wisconsin Alumni Research Foundation, Dr. DeLuca discusses the fascinating history of the discovery and applications of Vitamin D. Also touched upon is the creation of the Wisconsin Alumni Research Foundation, whose mission is to transform university research into real products that benefit the society at large.
Stem Cell Primer

Stem cells do not just come from human embryo, but also found in most body and extra embryonic tissues and membranes. What separates stem cells from other cells is its ability to self renew or make more of them without ageing. Normally when cell divides the DNA loses its end tips known as the telomeres,and this thought to be what makes cell age, these things doesn't happen in stem cells, but still retain the same length. this is one thing that no other body cell can do.
Systemic and Pulmonary Circulation
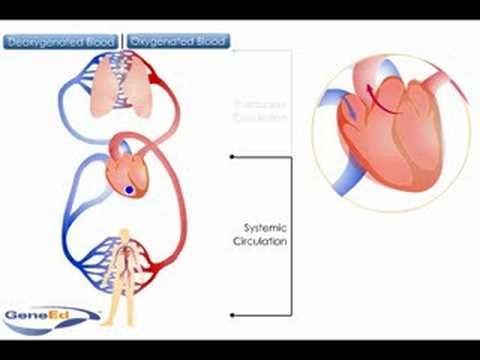
The circulatory system can be viewed as two circuits - the pulmonary circuit and systemic circuit.
Pulmonary circulation involves the transport of blood to and from the lungs. Deoxygenated blood returns to the right side of the heart. This blood is then pumped by the right ventricle into the pulmonary artery (deoxygenated blood) and into the capillaries of the lungs. In the capillaries of the alveoli, oxygen enters the blood and binds to hemoglobin in red blood cells as carbon dioxide diffuses out of the blood into the lungs for removal. The oxygenated blood then travels in the pulmonary vein back to the left atrium of the heart where it joins the systemic circuit.
Systemic circulation involves the transport of blood to and from all the tissues of the body. This circuit is much larger than the pulmonary circuit and so the walls of the left ventricle of the heart are much larger than on the right side. This thicker muscle generates the force required to pump blood all around the systemic circuit. Oxygenated blood is pumped by the left ventricle into the aorta. The aorta then branches into many smaller arteries that carry blood to all areas of the body. In the capillaries, oxygen is delivered to cells and carbon dioxide is picked up for removal. The deoxygenated blood then returns to the anterior vena cava from the upper body and the posterior vena cava from the lower body. Both vessels enter the right atrium of the heart and blood is returned back to the pulmonary circuit for oxygenation.
Colonoscopy

Colonoscopy is the endoscopic examination of the large colon and the distal part of the small bowel with a CCD camera or a fiber optic camera on a flexible tube passed through the anus. It may provide a visual diagnosis (e.g. ulceration, polyps) and grants the opportunity for biopsy or removal of suspected lesions. Virtual colonoscopy, which uses 2D and 3D imagery reconstructed from computed tomography (CT) scans or from nuclear magnetic resonance (MR) scans, is also possible, as a totally non-invasive medical test, although it is not standard and still under investigation regarding its diagnostic abilities. Furthermore, virtual colonoscopy does not allow for therapeutic maneuvers such as polyp/tumor removal or biopsy nor visualization of lesions smaller than 5 millimeters.

 If a growth or polyp is detected using CT colonography, a standard colonoscopy would still need to be performed. Colonoscopy can remove polyps as small as one millimeter or less. Once polyps are removed, they can be studied with the aid of a microscope to determine if they are precancerous or not. Colonoscopy is similar to but not the same as sigmoidoscopy. The difference between colonoscopy and sigmoidoscopy is related to which parts of the colon each can examine. Sigmoidoscopy allows doctors to view only the final two feet of the colon, while colonoscopy allows an examination of the entire colon, which measures four to five feet in length. Often a sigmoidoscopy is used as a screening procedure for a full colonoscopy. In many instances a sigmoidoscopy is performed in conjunction with a fecal occult blood test (FOBT), which can detect the formation of cancerous cells throughout the colon. Other times, a sigmoidoscopy is preferred to a full colonoscopy in patients having an active flare of ulcerative colitis or Crohn's disease to avoid perforation of the colon. Additionally, surgeons have lately been using the term pouchoscopy to refer to a colonoscopy of the ileo-anal pouch.
If a growth or polyp is detected using CT colonography, a standard colonoscopy would still need to be performed. Colonoscopy can remove polyps as small as one millimeter or less. Once polyps are removed, they can be studied with the aid of a microscope to determine if they are precancerous or not. Colonoscopy is similar to but not the same as sigmoidoscopy. The difference between colonoscopy and sigmoidoscopy is related to which parts of the colon each can examine. Sigmoidoscopy allows doctors to view only the final two feet of the colon, while colonoscopy allows an examination of the entire colon, which measures four to five feet in length. Often a sigmoidoscopy is used as a screening procedure for a full colonoscopy. In many instances a sigmoidoscopy is performed in conjunction with a fecal occult blood test (FOBT), which can detect the formation of cancerous cells throughout the colon. Other times, a sigmoidoscopy is preferred to a full colonoscopy in patients having an active flare of ulcerative colitis or Crohn's disease to avoid perforation of the colon. Additionally, surgeons have lately been using the term pouchoscopy to refer to a colonoscopy of the ileo-anal pouch.
Vitamin D Deficiency
Can vitamin D In this Lecture Robert P. Heaney, M.D., F.A.C.P., F.A.C.N.speaks about Vitamin How vitamin D can help prevent certain cancers and other diseases such as type 1 diabetes, cardiovascular disease, and certain autoimmune and chronic diseases.
About speaker
Dr. Heaney is an internationally recognized expert in the field of bone biology and calcium nutrition. He has worked for over 45 years in the study of osteoporosis and calcium physiology, and has published more than 300 original papers, chapters, monographs, and reviews in scientific and educational fields. He is a frequently invited editorial writer for the major medical journals, including JAMA, the American Journal of Clinical Nutrition, Annals of Internal Medicine, Clinical Chemistry, Metabolism, and the Journal of Clinical Endocrinology and Metabolism. He serves on numerous nutrition industry scientific advisory panels. Dr. Heaney was a member of the Panel on Calcium and Related Nutrients involved in the development of the new Dietary Reference Intakes for these nutrients, and his laboratory has done bioavailability testing for many of the major food companies in North America. His professional honors and awards include two that recognize his research in calcium physiology and calcium requirements in humans: honorary membership in the American Dietetic Association (1990) and election to Fellowship in the American College of Nutrition (1993) He is also the recipient of the Frederic C. Bartter Award of the American Society for Bone and Mineral Research (1994) and the Best Scientific Paper Award of the American College of Nutrition (2001). He has received three awards in 2003: the Scientific Prize of the Institut Candia (France) for his contributions to raising awareness of calcium and its health benefits; the E.V. McCollum Award of the American Society for Clinical Nutrition in recognition of his contributions to nutritional science and medicine; and the McCollum International Lectureship of the American Society of Nutritional Sciences.
Allergy

Allergy is a disorder of the immune system often also referred to as atopy. Allergic reactions occur to environmental substances known as allergens; these reactions are acquired, predictable and rapid. Strictly, allergy is one of four forms of hypersensitivity and is called type I (or immediate) hypersensitivity. It is characterized by excessive activation of certain white blood cells called mast cells and basophils by a type of antibody known as IgE, resulting in an extreme inflammatory response. Common allergic reactions include eczema, hives, hay fever, asthma, food allergies, and reactions to the venom of stinging insects such as wasps and bees.
In the early stages of allergy, a type I hypersensitivity reaction against an allergen, encountered for the first time, causes a response in a type of immune cell called a TH2 lymphocyte, which belongs to a subset of T cells that produce a cytokine called interleukin-4 (IL-4). These TH2 cells interact with other lymphocytes called B cells, whose role is production of antibodies. Coupled with signals provided by IL-4, this interaction stimulates the B cell to begin production of a large amount of a particular type of antibody known as IgE. Secreted IgE circulates in the blood and binds to an IgE-specific receptor (a kind of Fc receptor called FcεRI) on the surface of other kinds of immune cells called mast cells and basophils, which are both involved in the acute inflammatory response. The IgE-coated cells, at this stage are sensitized to the allergen.
Menopause

Menopause starts as the functioning of the ovaries begin to change. The ripening and release of the ovum (which, during the reproductive years leads to ovulation and then menstruation if pregnancy does not occur), becomes unpredictable. Ovulations start to be skipped, and the menstrual cycle starts to become less reliable in timing. As these changes become more pronounced, periods start to be skipped, and other perimenopausal symptoms may appear.
After a number of years of erratic functioning, the ovaries almost completely stop producing progesterone and two out of the three estrogen hormones: estradiol and estriol. Estrone is one estrogen which is still produced in reasonable amounts in post-menopausal women. Testosterone levels decrease; however, a decrease in testosterone levels begins gradually in young adulthood. Testosterone levels are thought not to drop significantly during the menopause transition because the stroma of the postmenopausal ovary and the adrenal gland still continue to secrete small amounts of testosterone, even during post-menopause.
Menopause is the end of the reproductive years rather than the beginning, and thus it is the opposite of menarche, nonetheless it can usefully be compared with that event: the menopause transition years are in many ways similar to puberty in that women experience hormonal fluctuations which usher in a new stage of life. Similar to pubescent girls, menopausal women are often unfamiliar or uneducated about the types of bodily changes they are undergoing, and may require time to adjust to the accompanying effects of these changes.
Subscribe to:
Posts (Atom)




